Bruce
Deagle, Post-Doc
email: bdeagle@uvic.ca
The focus of my research has been using information in DNA
to address ecological and evolutionary questions. I am currently working on a stickleback
population genomics project in Tom Reimchen’s laboratory; we are working
in close collaboration with David Kingsley’s group
at
Postdoctoral
work: SNP variation in Haida Gwaii threespine stickleback
Studies of Haida Gwaii stickleback
over the last 35 years have revealed a remarkable level of phenotypic variation
within and among freshwater populations, providing a classic example of
adaptive radiation (see Reimchen publications).
With
recent completion of a high-quality reference genome sequence, the threespine
stickleback is also quickly becoming an important model in the field of
evolutionary genomics (see
Kingsley publications). In this postdoctoral project we
are obtaining a genomic perspective of the genetic structure and history of Haida
Gwaii stickleback using a genome-wide SNP (single nucleotide polymorphisms)
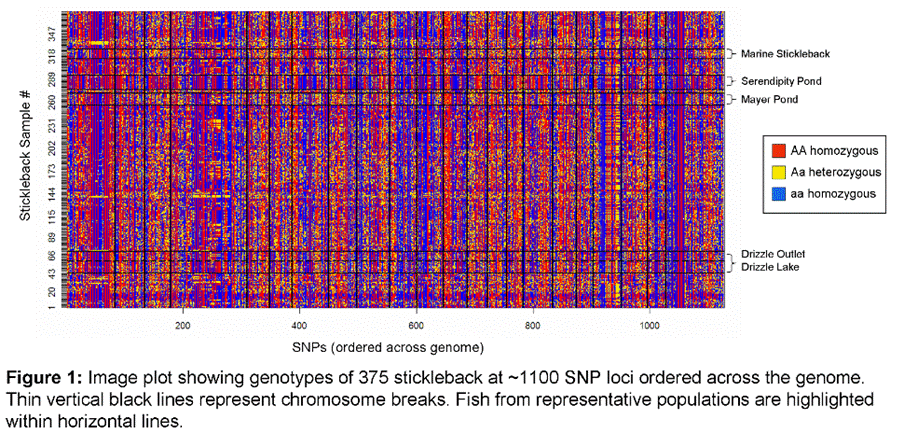
genotyping system (Illumina® GoldenGate) developed at Stanford (Figure
1). Particular areas of interest include:
(A) Phylogeography of the Haida Gwaii freshwater populations
(B) The population genomics of
parapatric stream/lake pairs of stickleback
(C) Associations between genetic markers and particular morphological
traits
(A)
Phylogeography
Since phylogenetic hypotheses
underlie most conclusions in studies of adaptive evolution, a robust
phylogeographic study in this model system would be invaluable. Initial
analysis shows that populations from within a watershed cluster together and
separate watersheds are generally independent. A few cases of genetic linkages
between separate watersheds in adjoining geographical areas may reflect past
watershed connections, or possibly ongoing gene flow. Populations that do not group with others are either: a single
sampling site within a watershed, close to a river mouth (continued marine gene
flow), or are isolated lakes with low heterozygosity which obscures historical
relationships.
(B) Parapatric
stream/lake pairs of stickleback
Parapatric
lake and stream populations of stickleback are a classic example of ecological
speciation. Two watersheds on Haida Gwaii (Drizzle and Mayer) contain
previously described stream/lake pairs (Moodie 1972 Can. J. Zool. 50: 721-732; Reimchen et al. 1985 Can.
J. Zool. 63:294-2951). To investigate the genetics underlying stream/lake
divergence we SNP genotyped fish from (1)
Data
confirm independent origin of these stream/lake pairs in the separate
watersheds (Figure 2 ). Within the Mayer watershed, fish
from two inlet creeks are closer to each other than to
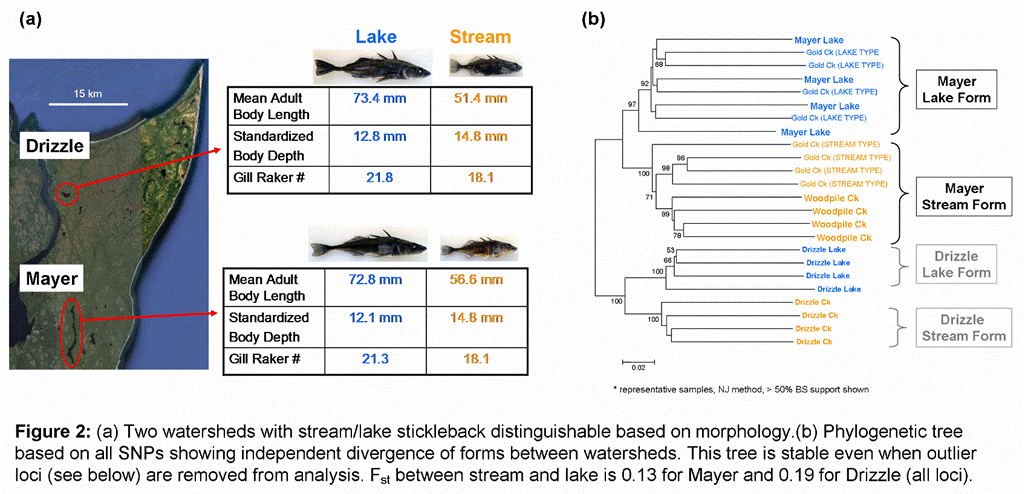
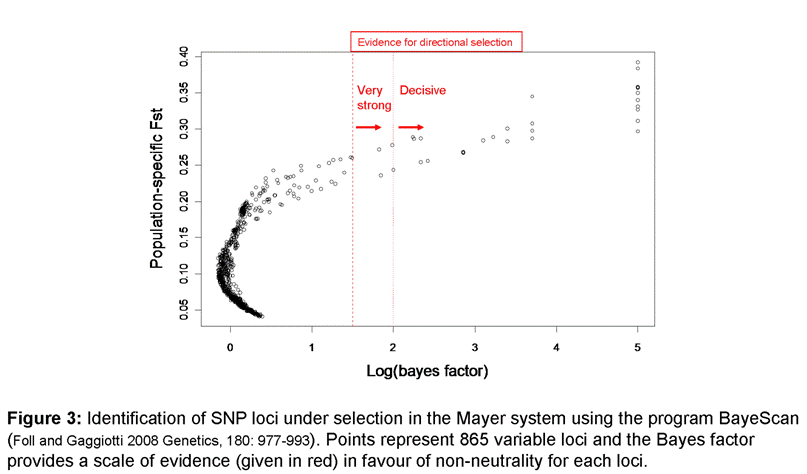
Genomic
divergence between stream and lake forms within a watershed was highly
heterogeneous: most loci show moderate levels of divergence but some loci
(outliers) were highly differentiated and are likely linked to genes under
strong divergent selection (Figure 3).
(C) Associations between
SNPs and morphological traits
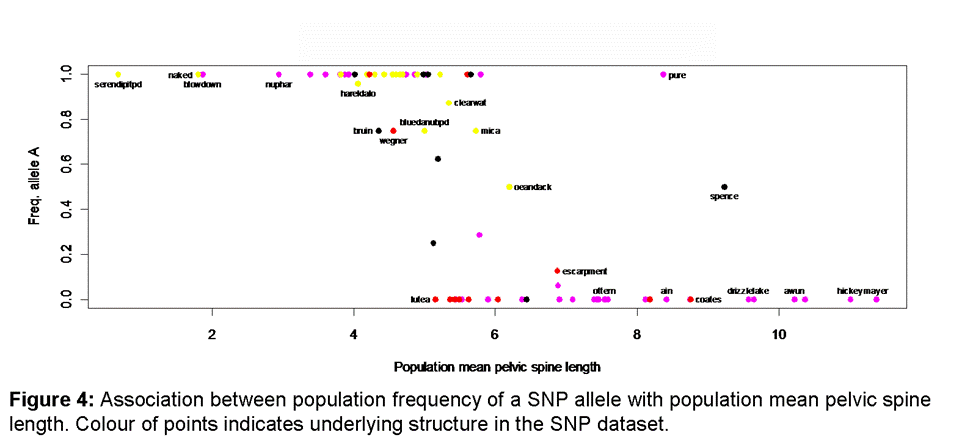
Populations
of Haida Gwaii stickleback exhibit a range of phenotypes defined by the location-specific
selective landscapes (Moodie
and Reimchen 1976 Systematic Zoology 25: 49-61). With morphological data from
over 100 populations (Reimchen
et al. In prep) and SNP data that characterizes
individuals from these populations, it may be possible to identify loci linked
to particular morphological traits (Figure 4).
PhD work at the University of Tasmania:
DNA-based methods to study diet
Studying animal diet is a difficult undertaking especially
in the marine environment where it is rarely possible to observe feeding
activity. Some prey remains survive digestion and morphological analysis of
remains in faecal material can provide an indication of what predators are
eating. In my PhD I looked at the potential for use of DNA-based identification
methods in dietary studies of marine predators. When I started it was clear
that prey DNA was present in animal faeces but how useful it would be for
dietary studies was an open question. So, in collaboration with researchers
from the Marine Mammal Unit at UBC,
I started by looking at samples collected from captive Steller
sea lions to address some basic questions:
·
 Can prey DNA be reliably detected in the soft matrix of faecal
samples?
Can prey DNA be reliably detected in the soft matrix of faecal
samples?
·
Can DNA from prey items fed as a small
proportion of the diet be detected?
·
Are the relative amounts of DNA
recovered from prey species proportional to their mass in the diet?
·
What is the quality of the prey
DNA recovered?
These questions
are the focus of three papers from my thesis:
Deagle
BE, Tollit DJ, Jarman SN, Hindell MA, Trites AW, Gales NJ (2005) Molecular
scatology as a tool to study diet: analysis of prey DNA in scats from captive
Steller sea lions. Molecular Ecology 14: 1831–1842. (pdf)
Deagle
BE, Eveson JP, Jarman SN (2006) Quantification of damage in DNA recovered from
highly degraded samples — a case study on DNA in faeces. Frontiers in
Zoology e3:11. (pdf)
Deagle
BE, Tollit DJ (2007) Quantitative analysis of prey DNA in pinniped faeces:
potential to estimate diet composition? Conservation Genetics 8: 743–747.
(pdf)
The
methods were further evaluated in a field-based study looking at the diet of
macaroni penguins at Heard Island.
Most recently I have been involved in projects using high-throughput sequencing
to look at diet of Australian fur seals and little penguins:
Deagle
BE, Gales NJ, Evans K, Jarman SN, Robinson S, Trebilco R, Hindell MA (2007)
Studying seabird diet through genetic analysis of faeces: a case study on
macaroni penguins (Eudyptes chrysolophus).
PLoS ONE 2: e831. (pdf)
Deagle
BE, Kirkwood R, Jarman
SN (2009) Analysis of Australian fur seal diet by pyrosequencing prey DNA in
faeces. Molecular Ecology 18: 2022–2038.
Deagle BE, Chiaradia A, McInnes J,
Jarman SN (2010) Pyrosequencing
faecal DNA to determine diet of little penguins: is what goes in what comes
out? Conservation
Genetics 11: 2039–2048.


For my other papers, please see publication list.





